-Search query
-Search result
Showing all 47 items for (author: schwab & ra)

EMDB-16128: 
Tomogram of a late endosome of A549 cell infected with influenza A virus.
Method: electron tomography / : Klein S, Chlanda P

EMDB-16129: 
Tomogram of a late endosome of A549 cell infected with influenza A virus (Figure 6C).
Method: electron tomography / : Klein S, Chlanda P

EMDB-16130: 
Tomogram of a late endosome of A549 cell infected with influenza A virus (Figure 6E)
Method: electron tomography / : Klein S, Chlanda P

EMDB-16131: 
Tomogram of a late endosome of A549 cell infected with influenza A virus (Figure 6G)
Method: electron tomography / : Klein S, Chlanda P

EMDB-16132: 
Tomogram of a late endosome of A549 cell infected with influenza A virus (Figure 6I,K,M)
Method: electron tomography / : Klein S, Chlanda P

EMDB-16133: 
Tomogram of a late endosome of A549 cell infected with influenza A virus (Figure 6O)
Method: electron tomography / : Klein S, Chlanda P

EMDB-15705: 
Tomogram of a late endosome of A549 cell (Figure 1R)
Method: electron tomography / : Klein S, Chlanda P

EMDB-15707: 
Tomogram of a late endosome of A549 cell treated with IFN-beta (Figure 1T)
Method: electron tomography / : Klein S, Chlanda P
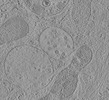
EMDB-15708: 
Tomogram of a late endosome of A549-IFITM3 cell (Figure 1V)
Method: electron tomography / : Klein S, Chlanda P

EMDB-15131: 
Tomogram of a late endosome of A549-IFITM3 cells infected with influenza A virus (Figure S7)
Method: electron tomography / : Klein S, Chlanda P
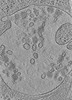
EMDB-15130: 
Tomogram of a late endosome of A549-IFITM3 cells infected with influenza A virus (Figure 4E)
Method: electron tomography / : Klein S, Chlanda P
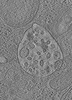
EMDB-15132: 
Tomogram of a late endosome of A549-IFITM3 cells infected with influenza A virus (Figure S8)
Method: electron tomography / : Klein S, Chlanda P

EMDB-15133: 
Tomogram of a late endosome of A549-IFITM3 cells infected with influenza A virus (Figure S9)
Method: electron tomography / : Klein S, Chlanda P

EMDB-13764: 
Structure of Hedgehog acyltransferase (HHAT) in complex with megabody 177 bound to non-hydrolysable palmitoyl-CoA (Composite Map)
Method: single particle / : Coupland C, Carrique L, Siebold C

PDB-7q1u: 
Structure of Hedgehog acyltransferase (HHAT) in complex with megabody 177 bound to non-hydrolysable palmitoyl-CoA (Composite Map)
Method: single particle / : Coupland C, Carrique L, Siebold C

EMDB-14578: 
Structure of Hedgehog acyltransferase (HHAT) in complex with megabody 177 bound to non-hydrolysable palmitoyl-CoA (Consensus Map)
Method: single particle / : Coupland C, Carrique L, Siebold C

EMDB-13860: 
Structure of Hedgehog acyltransferase (HHAT) in complex with megabody 177 bound to IMP-1575
Method: single particle / : Coupland C, Carrique L, Siebold C

PDB-7q6z: 
Structure of Hedgehog acyltransferase (HHAT) in complex with megabody 177 bound to IMP-1575
Method: single particle / : Coupland C, Carrique L, Siebold C

EMDB-13841: 
Focused refinement of Hedgehog acyltransferase (HHAT) in complex with megabody 177 bound to non-hydrolysable palmitoyl-CoA
Method: single particle / : Coupland C, Carrique L, Siebold C

EMDB-13842: 
Focused refinement of Hedgehog acyltransferase (HHAT) in complex with megabody 177 - megabody core
Method: single particle / : Coupland C, Carrique L, Siebold C

EMDB-12940: 
SARS-CoV-2-Induced Reshaping of Subcellular Morphologies
Method: electron tomography / : Cortese M, Schwab Y, Bartenschlager R, Schorb M

EMDB-12286: 
Cryo-EM structure of the ternary complex between Netrin-1, Neogenin and Repulsive Guidance Molecule B
Method: single particle / : Robinson RA, Griffiths SC, van de Haar LL, Malinauskas T, van Battum EY, Zelina P, Schwab RA, Karia D, Malinauskaite L, Brignani S, van den Munkhof M, Dudukcu O, De Ruiter AA, Van den Heuvel DMA, Bishop B, Elegheert J, Aricescu AR, Pasterkamp RJ, Siebold C

PDB-7ndg: 
Cryo-EM structure of the ternary complex between Netrin-1, Neogenin and Repulsive Guidance Molecule B
Method: single particle / : Robinson RA, Griffiths SC, van de Haar LL, Malinauskas T, van Battum EY, Zelina P, Schwab RA, Karia D, Malinauskaite L, Brignani S, van den Munkhof M, Dudukcu O, De Ruiter AA, Van den Heuvel DMA, Bishop B, Elegheert J, Aricescu AR, Pasterkamp RJ, Siebold C

EMDB-11938: 
Ternary complex of full-length Caspase-8 and FADD
Method: single particle / : Fox JL, Ragan TJ, Dinsdale D, Fairall L, Schwabe JWR, Morone N, Cain K, MacFarlane M

EMDB-11939: 
Central region of Caspase-8:FADD ternary complex
Method: single particle / : Fox JL, Ragan TJ, Dinsdale D, Fairall L, Schwabe JWR, Morone N, Cain K, MacFarlane M

EMDB-11940: 
Ternary complex of Full length Caspase-8 with FADD and FLIPs
Method: single particle / : Fox JL, Ragan TJ, Dinsdale D, Fairall L, Schwabe JWR, Morone N, Cain K, MacFarlane M

EMDB-11941: 
CryoEM analysis of ternary complex of full-length Caspase-8 with FADD and FLIPs
Method: single particle / : Fox JL, Ragan TJ, Dinsdale D, Fairall L, Schwabe JWR, Morone N, Cain K, MacFarlane M

EMDB-11837: 
Structure of the MTA1/HDAC1/MBD2 NURD deacetylase complex
Method: single particle / : Millard CJ, Fairall L

EMDB-11838: 
Structure of the core MTA1/HDAC1/MBD2 NURD deacetylase complex
Method: single particle / : Millard CJ, Fairall L

EMDB-11839: 
Structure of the extended MTA1/HDAC1/MBD2/RBBP4 NURD deacetylase complex
Method: single particle / : Millard CJ, Fairall L

PDB-7ao8: 
Structure of the MTA1/HDAC1/MBD2 NURD deacetylase complex
Method: single particle / : Millard CJ, Fairall L, Ragan TJ, Savva CG, Schwabe JWR

PDB-7ao9: 
Structure of the core MTA1/HDAC1/MBD2 NURD deacetylase complex
Method: single particle / : Millard CJ, Fairall L, Ragan TJ, Savva CG, Schwabe JWR

PDB-7aoa: 
Structure of the extended MTA1/HDAC1/MBD2/RBBP4 NURD deacetylase complex
Method: single particle / : Millard CJ, Fairall L, Ragan TJ, Savva CG, Schwabe JWR

EMDB-10660: 
Nup116_delta_NPC_25C
Method: subtomogram averaging / : Allegretti M, Zimmerli CE, Rantos V, Wilfling F, Ronchi P, Fung HKH, Lee CW, Hagen W, Turonova B, Karius K, Zhang X, Mulller C, Schwab Y, Mahamid J, Pfander B, Kosinski J, Beck M

EMDB-10661: 
Nup116delta_NPC_37C
Method: subtomogram averaging / : Allegretti M, Zimmerli CE, Rantos V, Wilfling F, Ronchi P, Fung HKH, Lee CW, Hagen W, Turonova B, Karius K, Zhang X, Muller C, Schwab Y, Mahamid J, Pfander B, Kosinski J, Beck M

EMDB-10198: 
In-cell S cerevisiae nuclear pore complex
Method: subtomogram averaging / : Beck M, Allegretti M

EMDB-11041: 
The structure of the dimeric HDAC1/MIDEAS/DNTTIP1 MiDAC deacetylase complex
Method: single particle / : Fairall L, Saleh A, Ragan TJ, Millard CJ, Savva CG, Schwabe JWR

EMDB-11042: 
The structure of the tetrameric HDAC1/MIDEAS/DNTTIP1 MiDAC deacetylase complex
Method: single particle / : Fairall L, Saleh A, Ragan TJ, Millard CJ, Savva CG, Schwabe JWR

PDB-6z2j: 
The structure of the dimeric HDAC1/MIDEAS/DNTTIP1 MiDAC deacetylase complex
Method: single particle / : Fairall L, Saleh A, Ragan TJ, Millard CJ, Savva CG, Schwabe JWR

PDB-6z2k: 
The structure of the tetrameric HDAC1/MIDEAS/DNTTIP1 MiDAC deacetylase complex
Method: single particle / : Fairall L, Saleh A, Ragan TJ, Millard CJ, Savva CG, Schwabe JWR

EMDB-10626: 
Negative stain map of CoREST complex (LSD1:RCOR1:HDAC1)
Method: single particle / : Song Y, Fairall L, Ragan TJ, Savva CG, Saleh A, Morone N, Schwabe JWR

EMDB-10627: 
Cryo-EM map of glutaraldehye cross-linked CoREST complex (LSD1:RCOR1:HDAC1)
Method: single particle / : Song Y, Fairall L, Ragan TJ, Savva CG, Morone N, Schwabe JWR

EMDB-10628: 
Cryo EM map of BS3 crosslinked CoREST complex (LSD1:RCOR1:HDAC1)- closed form
Method: single particle / : Song Y, Fairall L, Ragan TJ, Savva CG, Morone N, Schwabe JWR

EMDB-10629: 
Cryo EM map of BS3 crosslinked CoREST complex (LSD1:RCOR1:HDAC1) - open form
Method: single particle / : Song Y, Fairall L, Ragan TJ, Savva CG, Morone N, Schwabe JWR

EMDB-10630: 
Interaction of the CoREST complex with a nucleosome with 185 bp 601 sequence DNA and a propargylamine mimic of dimethy Lys4 histone H3
Method: single particle / : Song Y, Fairall L, Wu M, Ragan TJ, Savva CG, Morone N, Cole PA, Schwabe JWR

PDB-6rvd: 
Revised cryo-EM structure of the human 2:1 Ptch1-Shh complex
Method: single particle / : El Omari K, Rudolf AF, Kowatsch C, Malinauskas T, Kinnebrew M, Ansell TB, Bishop B, Pardon E, Schwab RA, Qian M, Duman R, Covey DF, Steyaert J, Wagner A, Sansom MSP, Rohatgi R, Siebold C

EMDB-3399: 
Structure of the core NuRD complex (MTA1:HDAC1:RBBP4)
Method: single particle / : Millard CJ, Saleh A, Morris K, Fairall L, Smith CJ, Schwabe JWR
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
